Efficient evidence selection for systematic reviews in traditional Chinese medicine | BMC Medical Research Methodology

Overall description of the proposed clinical evidence selection system
This novel literature selection methodology comprised four steps:
Initial literature download
We started by retrieving a set of original publications from various electronic databases, employing search strategies tailored to align with the research topic.
Text segmentation and conversion
We employed machine vision technology to segment the downloaded publications into manageable blocks based on paragraph differences. These blocks were then converted into computer-readable text files (in TXT format) using Optical Character Recognition (OCR) with open-source tools. This process enabled us to process the full text for subsequent information extraction.
Evidence extraction
The Evi-BERT model, integrated with rule-based extraction methods, systematically retrieved structural evidence from various sections of the articles. This ensured that key clinical information was extracted accurately and efficiently.
Screening and selection
We applied Boolean logic to identify the targeted clinical elements. Logical judgment criteria were used to determine whether the literature met the specified inclusion or exclusion criteria, ultimately selecting the qualified RCTs.
Detailed workflow of the clinical evidence selection system
To validate our approach, we compared this new procedure to traditional manual methods by re-screening original research literature from ten high-quality systematic reviews (SR1-SR10) focused on TCM topics. These reviews, published over the last decade, cover a range of topics, including pediatric diarrhea (SR1) [35], diabetic peripheral neuropathy (DPN) (SR2) [36], chronic obstructive pulmonary disease stable pulmonary (COPD) (SR3) [37], simple obesity (SR4) [38], diabetic kidney disease (DKD)(SR5) [39], knee osteoarthritis (SR6-8) [40,41,42], chronic atrophic gastritis (SR9) [43], and type 2 diabetes insulin resistance (SR10) [44]. Figure 1 illustrates the operational process of applying the methodology to the ten selected examples.
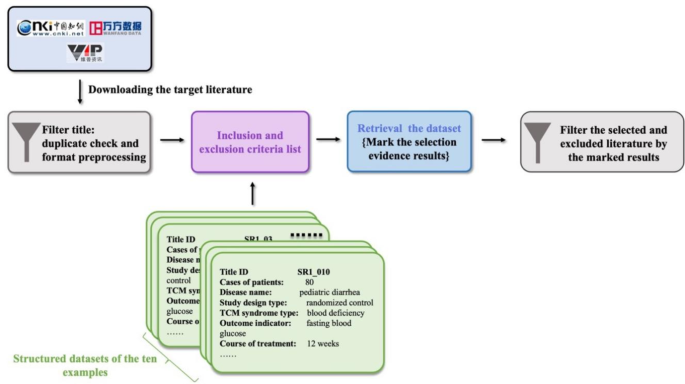
The concrete implementation framework of clinical evidence selection in the samples
Literature data collection and preprocessing
We began acquiring initial literature sets based on the search protocols used in the selected systematic reviews. Over 90% of the relevant literature was in Chinese, so the current method was designed specifically for Chinese-language articles. We searched literature from CNKI, VIP, WanFang and CBM. The retrieved literature was exported to a literature management tool, Note-express.
The publications, mostly in PDF or image formats, were then converted into editable TXT files using OCR technology with open-source tools such as PyMuPDF and OpenCV [33], as shown in Fig. 2. Given the standardized format of RCTs, we segmented the articles (TXT files) into 11 sections (Appendix 1) to facilitate efficient extraction of target information. Specific keywords and phrases were used to identify each section (Appendix 2), while irrelevant data, such as formatting errors (‘< p>’ and ‘\n’), were removed during a data-cleaning process.
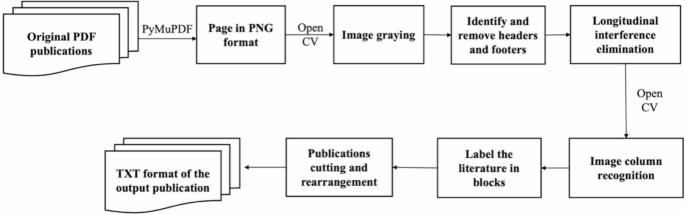
The original PDF publications’ identification process
Integrating inclusive and exclusive criteria
We merged the inclusion and exclusion criteria for the systematic reviews when they addressed the same issue. For example, if an inclusion criterion specified RCTs and the exclusion criterion ruled out uncontrolled studies, these were combined to simplify the screening process. Synonyms for key terms, such as “meta-analysis” and “text mining,” were also incorporated to enhance the accuracy of the automated exclusion process.
In cases where exclusion criteria could not be comprehensively covered by the code, corresponding inclusion criteria were created to maintain screening accuracy. This approach allowed us to filter out irrelevant studies without manual intervention, though certain complex cases still required human review (e.g., inability to calculate the 95% confidence interval; statistical method errors; and data errors). For instance, while it was not feasible to exclude all potential terms related to animal experiments, we established inclusion criteria focusing on key elements such as patients, randomization methods, and group names. Based on the fusion of inclusion and exclusion criteria, we summarized nine target information from eight sections for each systematic review (Table 2).
Evidence selection from literature
For evidence extraction, we used the Evi-BERT model combined with the precision-preferred comprehensive information extraction system. This allowed us to extract critical information across different sections of the articles, including titles, abstracts, and full texts.
The extraction system was designed to begin searching for relevant information in the most likely sections and gradually expand the search scope if the required data was not found. For example, when querying intervention methods, we started by searching the title and keywords, then extended the search to the abstract, followed by the study intervention in the methods section, and finally conducted a full-text search if no relevant information was extracted from the previous sections. This approach minimized the risk of including background information unrelated to the research question.
Three possible outcomes were labeled during the extraction process:
Extracted: Critical information was successfully retrieved.
Required for Manual Review: Potentially related information was flagged for further human verification.
Missed: No relevant information was found.
The extraction process continued to be refined until all ” Required for Manual Review” labeled results were resolved, ensuring accurate evidence extraction for each systematic review. Researchers carefully proofread any outcomes marked as “Required for Manual Review.” Additionally, to reduce potential errors, researchers also reviewed the information labeled as “Missed”, refining the rule-based components of the code as needed. This optimization process continued until no information was marked as “Required for Manual Review.” For each systematic review, a format matrix was created, cataloging the evidence elements from all relevant articles, along with labels indicating the reasons for inclusion or exclusion.
As shown in Fig. 3, the extraction progress allowed us to obtain the clinical text and numerical information necessary for literature filtering. Based on the inclusion and exclusion criteria, we developed a series of code blocks incorporating element dictionaries and Boolean operation patterns, to facilitate literature selection (Appendix 3). Clinical researchers can further optimize the code structure to suit various screening objectives.
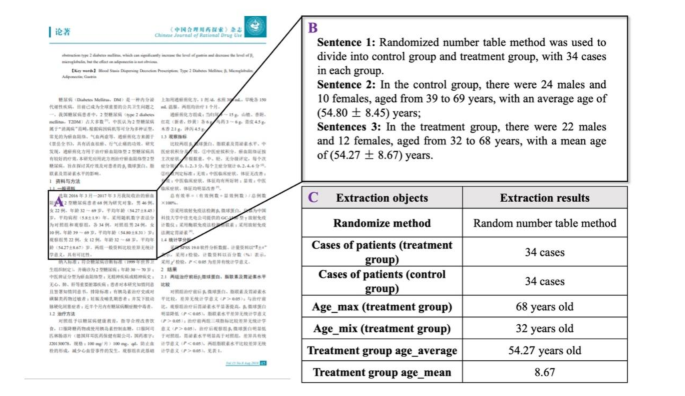
An example of paragraph searching, sentence splitting, and evidence extraction from electronic literature using Evi-BERT combined with rule-based extraction. (A) Identify the paragraph most relevant to the target information within the designated sections. (B) Extract the sentence containing the target number from the paragraph, such as: “In the treatment group, there are 22 males and 12 females, aged from 32 to 68 years, with a mean age of (54.27 ± 8.67) years.” (C) Match the phrases containing numbers, such as “32 to 68 years,” and extract the specific numbers, like “32” and “68,” to generate a format matrix
Excluded by Screening Program: Articles were automatically excluded if they contained designated exclusion terms or lacked required inclusion terms in titles, keywords, abstracts, or full texts. Numerical data, such as treatment durations, were also screened based on predefined criteria. For example, SR1 specifies that the intervention in the treatment group must last at least eight weeks. To meet this inclusion criterion, we extracted phrases such as “eight courses of treatment, seven days per course,” “a total of fifty-six days of treatment,” or “the course of treatment for both groups was eight weeks,” and convert them into “fifty-six days” for logical evaluation in the “course of treatment” field. If an article contains extracted data that does not meet the inclusion criteria or satisfies the exclusion criteria, it is labeled as “Excluded by Screen Program.”
Excluded by authors: Articles that could not be excluded through automated screening but did not meet the inclusion criteria or satisfied the exclusion criteria after manual review were labeled as excluded by authors.
Included: All remaining articles that passed the screening process were labeled as included.
link





