Experimental instruments
Ultra-high-performance liquid chromatograph (LC-30 A, Shimadzu), Mass spectrometer (TripleTOF 6600+, SCIEX), Centrifuge (5424R, Eppendorf), Constant temperature metal shaker (MU-G02-0448, Hangzhou MIO Instrument Co., Ltd.), analytical balance with a resolution of 0.0001 g (MS105DM, Mettler Toledo Instruments Co., Ltd.), centrifugal concentrator (CentriVap, LABCONCO), vortex mixer (VORTEX-5, Kyllin-Bell), ultrasonic cleaner (KQ5200E, Kunshan Instrument Co., Ltd.), Mouse Macrophages RAW 264.7 ( Shanghai Institutes for Life Sciences, Chinese Academy of Sciences).
Experimental materials
Methanol (HPLC grade, Merck, Cat. No.: 1.06007.4008), acetonitrile (HPLC grade, Merck, Cat. No.: 1.00030.4008), formic acid (HPLC grade, Aladdin, Cat. No.: 695076-100ML), anhydrous ethanol (Xilong Chemical, China).
Collection and screening of P. rubra components
Preparation of test samples
The P. rubra samples used in the experiment were collected from Hainan Province, China. Species identification was carried out by Mr. Li Rongtao, one of the senior researchers from the Hainan Branch of the Chinese Academy of Medical Sciences Institute of Medicinal Plants. Voucher specimen was prepared and deposited at the Hainan Provincial Chinese Medicine Herbarium under the collection number JJ20240915001. A total of 2 g of P. rubra was added into 50 mL of distilled water, which was then boiled in a water bath with reflux for 2 h. After cooling, the solution was filtered and transferred to a pre-dried, constant-weight evaporating dish. The leftover was cooled in a desiccator for 30 min and was quickly weighed. The residue was re-dissolved with a solvent to prepare a solution with a concentration of 1 mg/mL. A total of 100 µL of the solution was transferred into a 1.5 mL centrifuge tube, vortexed for 15 min, and centrifuged at 12,000 rpm at 4 °C for 3 min. The supernatant was filtered through a microporous membrane (0.22 μm pore size) and stored in an autosampler vial for subsequent LC-MS/MS analysis.
Chromatography-mass spectrometry collection conditions
The Waters ACQUITY UPLC HSS T3 Column (1.8 μm, 2.1 mm*100 mm) was selected, with 0.1% formic acid in water as mobile phase A and 0.1% formic acid in acetonitrile as mobile phase B. The column temperature was set to 40 °C, and the flow rate was 0.4 mL/min, with an injection volume of 4 µL. The gradient program started with 95% mobile phase A. At the fifth minute, the proportion of mobile phase A was reduced to 35%, then to 1% at the sixth minute, and maintained at 1% for 1.5 min. At the seventh minute and thirty-sixth second, the proportion of mobile phase A was restored to 95% and maintained until the end of the run at 10 min.
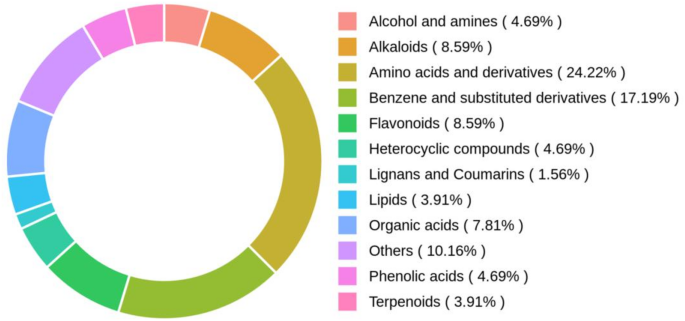
Screening of chemical components of P. rubra
The retention times of the chemical components detected in P. rubra were corrected, and peaks with a missing rate > 50% in the samples were filtered out. Metabolite identification was performed by searching and integrating public databases, prediction libraries, and the metDNA method. Finally, substances with an identification score above 0.5 and a QC sample CV value < 0.3 were extracted, followed by merging positive and negative modes (retaining the compounds with the highest qualitative level and smallest CV value). The SMILES numbers of the active compounds were retrieved using the PubChem database ( accessed on 1 November 2024), and preliminary screening was conducted using the SwissADME database ( accessed on 1 November 2024) with the following criteria: iLogP ≤ 5, nOHNH ≤ 5, nOH ≤ 10, OB ≥ 30%. Compounds not meeting these conditions were eliminated based on Lipinski’s five rules. Final screening was performed based on High GI absorption and Leadlikeness in Medicinal Chemistry, to confirm the effective active ingredients in P. rubra.
Prediction of active ingredient targets
The SwissTargetPrediction database ( accessed on November 1, 2024)1 and ChEMBL database ( accessed on November 1, 2024)2 were used to construct a target library for the components of P. rubra. The gene names of the target data were then standardized using the UniProt database ( accessed on 1 November 2024), and only human (Homo sapiens) target data were retained.
Construction of the target set related to respiratory tract infection diseases
Genes related to respiratory tract infection diseases were retrieved from the GeneCards ( accessed on November 1, 2024) and DisGeNet ( accessed on 1 November 2024) databases using “respiratory tract infection” as the keyword to search for the disease targets. Gene names were standardized using the UniProt database ( accessed on November 1, 2024)3. To ensure the reliability of the data, targets with a high correlation value (≥ 15.0) were selected.
PPI network construction and analysis
The gene targets regulated by P. rubra were intersected with the disease targets related to respiratory tract infections to identify the potential targets through which P. rubra exerts its effects in treating inflammation caused by respiratory infections. These potential targets were further screened to confirm the relevant targets for P. rubra’s therapeutic action in respiratory infection-induced inflammation. The resulting target gene set was then submitted to STRING ( accessed on 1 November 2024)4 to construct a PPI network, with Homo sapiens selected as the target species. A high-confidence interaction score threshold of 0.9 was applied, and independent targets were removed while adjusting the structure of the remaining targets. The resulting PPI network was visualized in Cytoscape v3.10.0, and the DEGREE values of all interacting targets were calculated and ranked using the Cytoscape v3.10.0 plugin. The core targets related to diseases caused by respiratory infection-induced inflammation were then selected for further analysis.
Gene ontology (GO) and KEGG pathway enrichment analysis
The potential targets of P. rubra in treating inflammation caused by respiratory infections were imported into the Metascape database ( accessed on 1 November 2024) and DAVID ( accessed on 1 November 2024) database for Gene Ontology (GO) and KEGG (www.kegg.jp/kegg/kegg1.html)5,6,7 pathway enrichment analyses. This could provide an abundant amount of biological information on the potential targets and further support the analysis on revealing the underlying mechanisms of action for P. rubra in treating inflammation.
Molecular docking
By combining the number of targets and core genes involved, active components and core targets were selected for docking to predict and obtain the binding energy of protein-ligand interactions. The compound structures in sdf format were obtained from the PubChem database, and the pdb files of the core target structures were retrieved from the RCSB PDB database ( accessed on 1 November 2024)8. Docking was performed using Autodock software (AutodockTool 1.5.7), and visualization was carried out with PyMOL (PyMol 2.5.0). The binding energy was used as an indicator to assess the binding activity and docking effect between the ligand and protein. A binding energy of < -1.4 kcal·mol-1 is generally considered to indicate a strong interaction between the two.
The effect of PRHT on the release of inflammatory cytokines in LPS-induced RAW264.7 cells
The RAW264.7 mouse macrophages were divided into three groups: blank group (with blank culture medium), control group (with culture medium without the drug), and drug concentration treatment group (with different concentrations of the drug). The cell viability was measured using the CCK-8 method by determining the absorbance. An LPS-induced RAW264.7 mouse cell inflammation model was also established, with the cells divided into the following groups: blank group (3 mL culture medium), model group (3 mL of LPS with a final concentration of 1 µg/mL), and treatment group (1 mL of LPS with a final concentration of 1 µg/mL + 2 mL of four different drug concentrations (0.025, 0.05, 0.1, 0.2 mg/mL)). After 24 h, the cell culture supernatants were collected, and the levels of TNF-α, IL-1β, and IL-6 were measured.
Log-phase cells were cultured in a 37 °C, 5% CO2 incubator for 24 h to allow for adaptation. The levels of TNF-α, NO, IL-1β, and IL-6 in the cell culture supernatant were measured using an enzyme-linked immunosorbent competitive assay (ELISA). The Griess method was used to determine the absorbance of the reaction products and calculate the NO content. The NaNO2 standard solutions were diluted in fresh cell culture medium (DMEM + 10% FBS) to concentrations of 0, 1, 2, 5, 10, 20, 40, 60, and 100 µmol/L. A total of 50 µL of the standard solutions or cell supernatant were added to a 96-well plate, with three replicates for each concentration. Griess Reagents I and II were pre-warmed to room temperature and then added to the plate. Upon color change, absorbance at 540 nm was immediately measured using a microplate reader.
$${\text{Cell}}\,{\text{viability}}\left( \% \right) = \left[ {\left( {{\text{Drug}}\,{\text{group}}\,{\text{OD}} – {\text{Blank}}\,{\text{group}}\,{\text{OD}}} \right)/\left( {{\text{Control}}\,{\text{group}}\,{\text{OD}} – {\text{Blank}}\,{\text{group}}\,{\text{OD}}} \right)} \right]*100$$
(1)
$${\text{Inhibition}}\,{\text{rate}}\left( \% \right) = \left[ {\left( {{\text{Control}}\,{\text{group}}\,{\text{OD}} – {\text{Drug}}\,{\text{group}}\,{\text{OD}}} \right)/\left( {{\text{Control}}\,{\text{group}}\,{\text{OD}} – {\text{Blank}}\,{\text{group}}\,{\text{OD}}} \right)} \right]*100$$
(2)
Statistical analysis method
The data were analyzed using appropriate statistical methods to evaluate the significance of the observed effects. Continuous variables were expressed as means ± standard deviation (SD). Group comparisons were performed using one-way analysis of variance (ANOVA), followed by Tukey’s post hoc test to identify specific differences between groups. In cases where data were not normally distributed, non-parametric tests. For molecular docking and gene expression analysis, binding energy scores and gene target interactions were assessed to determine the strength of the ligand-protein interactions. The results were considered statistically significant when the p-value was less than 0.05. To analyze the effects of PRHT on inflammatory cytokines in LPS-induced RAW264.7 cells, we performed an ANOVA with Tukey’s post hoc tests to compare cytokine release between the treatment and control groups. The significance level was set at p < 0.05 for all statistical tests. Data analysis was carried out using GraphPad Prism (version X) software.
link